
Open Access

Subscription Access
In silico identification of potent FDA approved drugs against Coronavirus COVID-19 main protease: A drug repurposing approach
Dhruv Kumar, Vaishali Chandel, Sibi Raj, Brijesh Rathi
Abstract
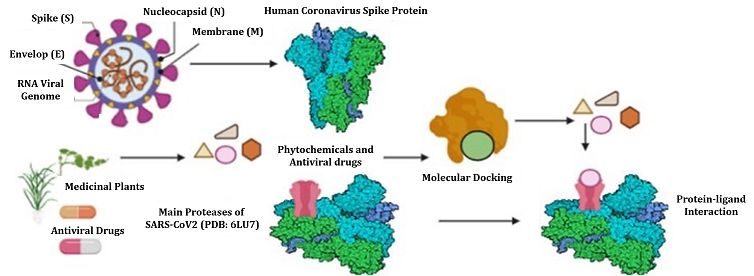
The Coronavirus COVID-19 Main Proteases play critical role in the propagation of the Novel Coronavirus (COVID-19). We have applied a bioinformatics approach of drug repurposing to identify possible potent inhibitors against Novel Coronavirus (COVID-19) through targeting COVID-19 Main Protease from FDA approved antiviral compounds and from the library of active phytochemicals. The compounds were screened using PyRx virtual screening tool. Total 19 best compounds were identified after screening, based on their highest binding affinity with respect to the other screened compounds. Out of 19, 6 best compounds were further screened based on their binding affinity and best ADME properties. Nelfinavir exhibited highest binding affinity -8.4 Kcal/mol and strong stability to interacted with the amino acid residues present on the active site of COVID-19 Main Protease. In addition to Nelfinavir (-8.4), Rhein (-8.1), Withanolide D (-7.8), Withaferin A (-7.7), Enoxacin (-7.4), and Aloe-emodin (-7.4) also showed good binding affinity and best ADME properties. Our findings suggest that these compounds can be used as potential inhibitors against COVID-19 Main Protease, which could be helpful in inhibiting the propagation of the Novel Coronavirus (COVID-19). Moreover, further investigation and validation of these inhibitors against Coronavirus would be very helpful to bring these molecules to the clinical settings.
Keywords
Novel Coronavirus; COVID-19; Protease; Molecular Docking; Drug Designing; ADME; Drug Repurposing
Full Text:
PDF

ISSN 2347–9825
Authors/visitors are advised to use Firefox browser for better experience of journal site.
Open Access: Researcher from developing/low economy countries can access the jorunal contents through WHO-HINARI .
 ISSN 2347-9825
ISSN 2347-9825

